DINO-3DRA
Leveraging 2D Foundation Model Semantics for 3D Cerebral Aneurysm Segmentation
We transfer frozen 2D VFM semantics into a 3D U-Net via slice-level embeddings, depth-coherence restoration, and calibrated residual fusion — achieving +13% Dice over nnU-Net with only 116K extra parameters and zero catastrophic failures on cross-dataset evaluation.
0.758
Aneurysm Dice+13%
over nnU-Net116K
Extra params0%
Failure rate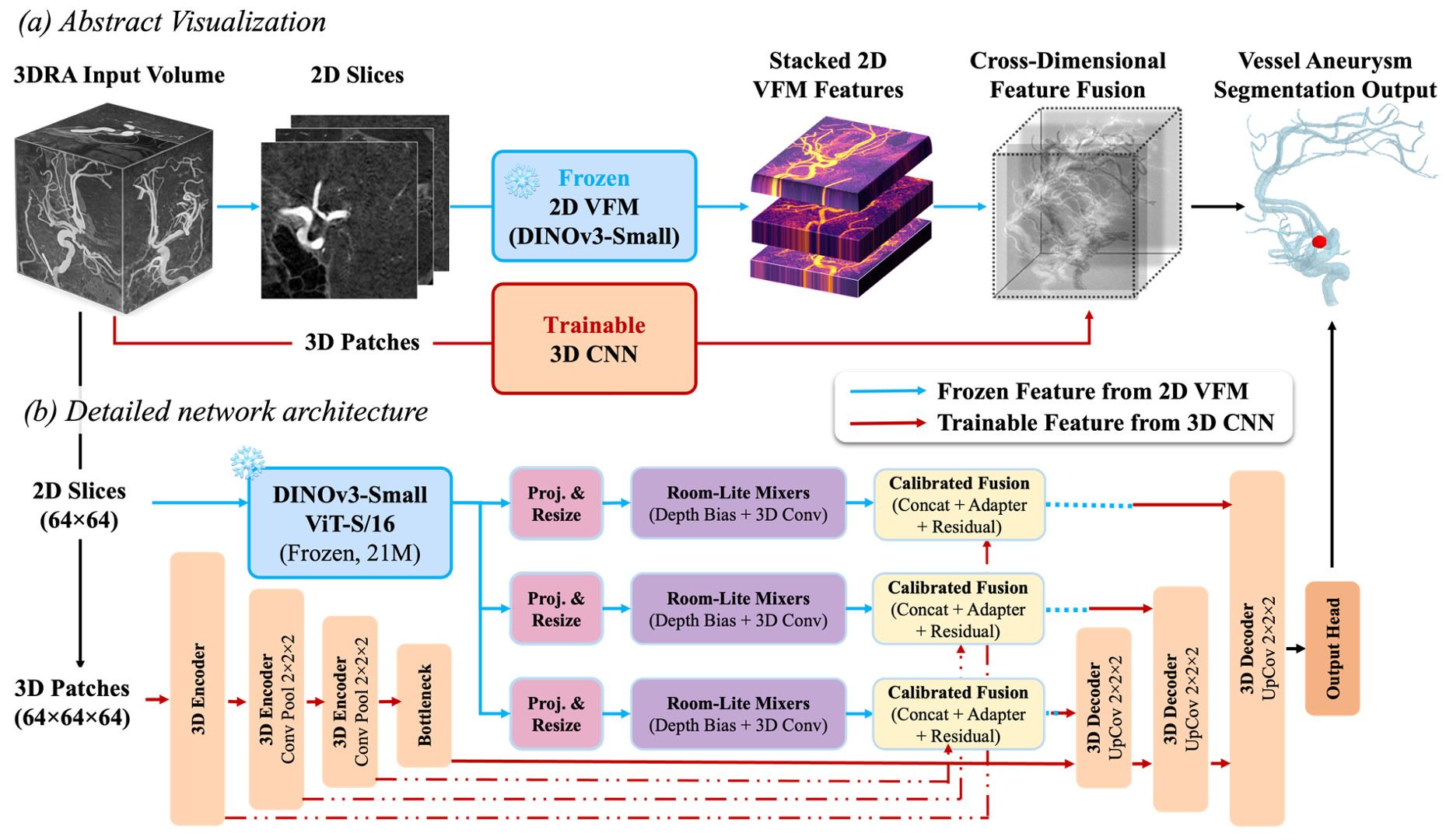
Abstract
Accurate aneurysm segmentation in 3D rotational angiography (3DRA) is hindered by extreme class imbalance, morphological similarity to vessels, and absent large-scale 3D pretraining. 2D vision foundation models encode dense structural priors from ~1.7 billion images, yet naïve slice-wise transfer fragments anatomical continuity and destabilises optimisation.
We propose DINO-3DRA, a dual-path framework achieving effective cross-dimensional semantic transfer by injecting frozen DINOv3 features into a 3D U-Net via Room-Lite spatial mixing and calibrated residual fusion. On multi-centre 3DRA data, DINO-3DRA achieves state-of-the-art aneurysm segmentation (Dice: 0.758; HD95: 2.75 mm; +13% over nnU-Net) with only 5.72M trainable parameters. Ablation studies confirm that gains arise from structured cross-dimensional transfer rather than loss design alone. Cross-dataset evaluation eliminates all catastrophic failure cases, demonstrating robust generalisation across heterogeneous imaging protocols.
The Challenge
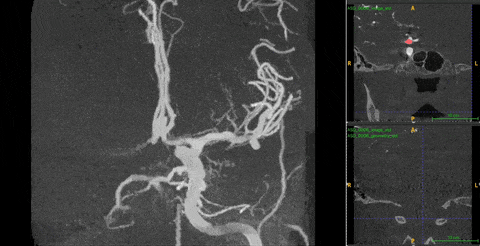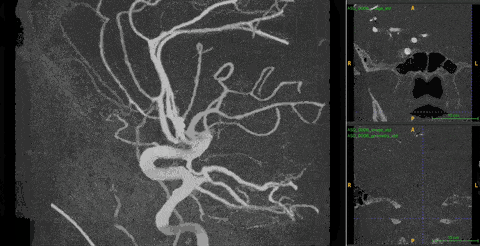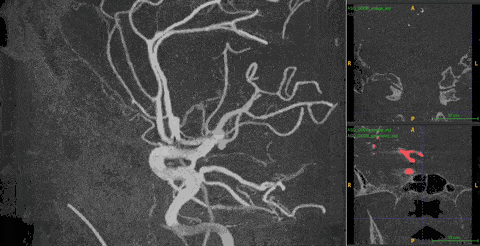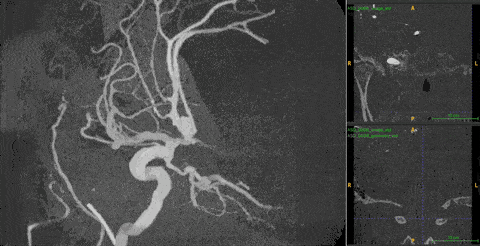
Ambiguity. 3DRA produces near-binary contrast — bright vessel lumen on dark background. On any single 2D slice, both a vessel cross-section and an aneurysm appear identical: a white circle on black. Distinguishing them requires observing how the cross-sectional shape changes across consecutive depth slices — a vessel maintains a consistent tube, while an aneurysm bulges and contracts.
The 2D–3D gap. The best 2D models (DINOv3) are trained on 1.7B images with rich structural priors. No equivalent large-scale 3D pretraining exists. But naïvely injecting 2D features into 3D models fails — it makes performance worse than using no 2D features at all.
Method
DINO-3DRA pairs a frozen 2D VFM with a trainable 3D U-Net. The frozen branch provides what kind of anatomy is in each slice. The trainable branch provides where exactly structures are. Two lightweight modules bridge the gap.
1 — Slice-Level Semantic Extraction. All 64 axial slices processed independently by frozen DINOv3-Small. 196 patch tokens globally averaged to a single 384-dim semantic embedding per slice. Projected via 1×1 convolutions and stacked into 3D volumes at three U-Net skip scales.
(B×64, 196, 384) → mean-pool → (B×64, 384) → (B, {24/48/96}, {64³/32³/16³})
2 — Room-Lite Depth Coherence. Learnable depth-positional bias B(z) + depthwise-separable 3D convolutions restore inter-slice continuity. Without this, Dice collapses −49%. 21K params total.
Y = X + GELU(GN(Conv₁ₓ₁(Conv₃ₓ₃(X + B(z)))))
3 — Calibrated Residual Fusion. GN-normalised DINO features injected as a learnable residual correction into U-Net skips. The residual bypass guarantees DINO can only help, never hurt.
F_fused = F_U + GELU(GN(W[F_U; GN(F_D)]))
Results
Ablation Study
Four of five DINO variants fall below the plain U-Net baseline. Only the full pipeline — Room-Lite + Calibrated Fusion — surpasses it.
| Aneurysm | Vessel | |||
|---|---|---|---|---|
| Configuration | Dice | HD95 | Dice | HD95 |
| Ours (Full) | 0.758 ± 0.234 | 2.75 | 0.897 ± 0.033 | 3.74 |
| Baseline U-Net | 0.711 ± 0.240 | 4.20 | 0.873 ± 0.046 | 5.41 |
| Random Features | 0.628 ± 0.279 | 6.15 | 0.885 ± 0.039 | 4.41 |
| DINOv2-small | 0.590 ± 0.252 | 6.50 | 0.868 ± 0.050 | 4.30 |
| w/o Calibration | 0.492 ± 0.301 | 10.12 | 0.857 ± 0.045 | 6.71 |
| w/o Room-Lite | 0.385 ± 0.310 | 22.08 | 0.857 ± 0.043 | 5.64 |
Loading visualization…
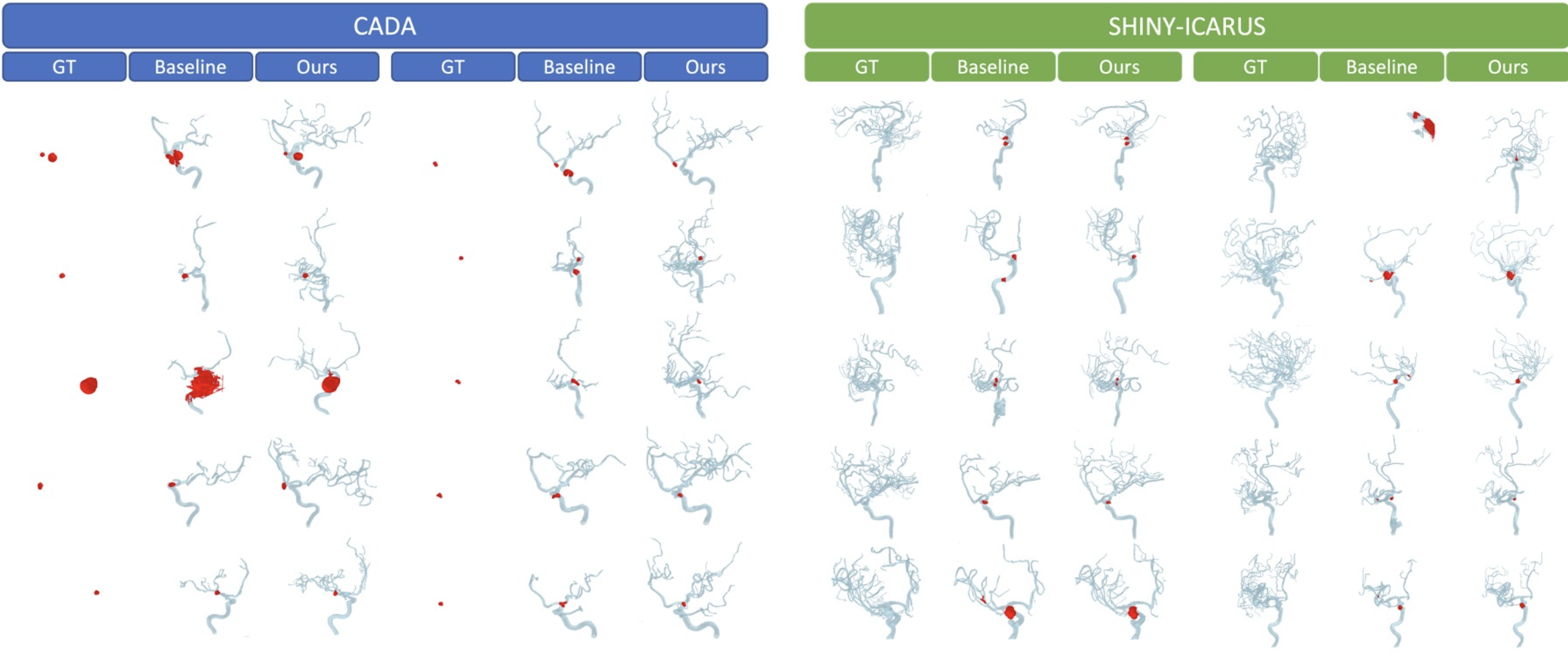
Cross-Dataset Generalisation
Evaluated on two external datasets without fine-tuning. DINO-3DRA eliminates all catastrophic failures (Dice < 0.5).
11.1% → 0%
Failures on CADA3.3% → 0%
Failures on SHINY-ICARUS75.8%
HD95 reduction (SI)| Dataset | Model | Dice | Jaccard | Vol. Sim. | HD95 | Failures |
|---|---|---|---|---|---|---|
| CADA | Baseline | 0.711 ± 0.186 | 0.577 ± 0.186 | 0.813 ± 0.128 | 5.16 | 11.1% |
| CADA | DINO-3DRA | 0.739 ± 0.093 | 0.594 ± 0.118 | 0.807 ± 0.109 | 3.58 | 0% |
| SHINY-ICARUS | Baseline | 0.718 ± 0.147 | 0.574 ± 0.129 | 0.808 ± 0.089 | 15.52 | 3.3% |
| SHINY-ICARUS | DINO-3DRA | 0.793 ± 0.049 | 0.660 ± 0.065 | 0.812 ± 0.056 | 3.76 | 0% |
BibTeX
@inproceedings{dino3dra2026,
title = {DINO-3DRA: Leveraging 2D Foundation Model
Semantics for 3D Cerebral Aneurysm Segmentation},
author = {Anonymized Authors},
booktitle = {International Conference on Medical Image Computing
and Computer-Assisted Intervention (MICCAI)},
year = {2026}
}